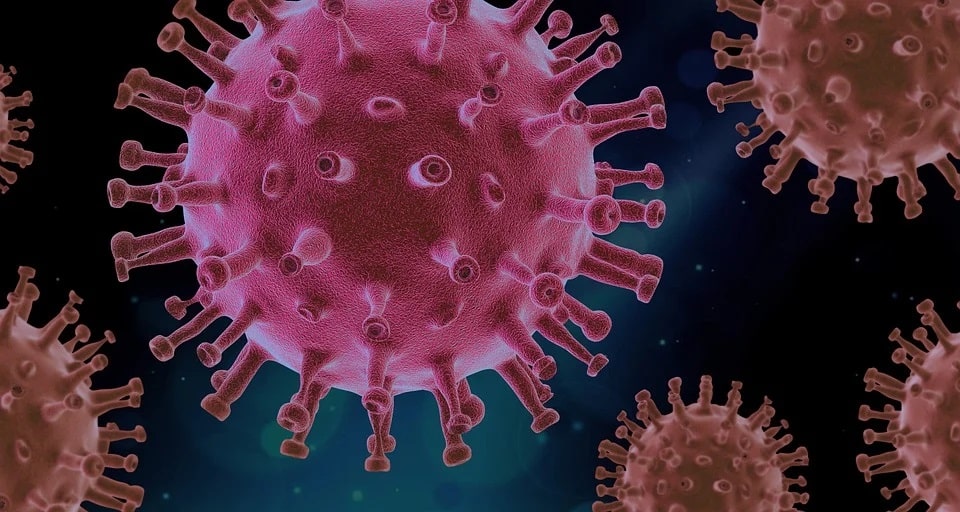
The virus that causes COVID-19, SARS-CoV-2, is a type of coronavirus, a large family of viruses named for the crown-like spikes on their surfaces. SARS-CoV-2 is a large RNA virus made up of around 30,000 nucleotides. A feature of RNA viruses is their relative inaccuracy in the replication process. Therefore, every new copy can present stochastic mutations. If the new mutation favours the virus over the host or has a competitive advantage over the other already circulating strains, it might become dominant. This is what is happening in the United Kingdom (UK) where a mutated virus first detected on September 20, 2020 has emerged and is already the most widespread form in the country.
At present, this new variant is referred to as “SARS-CoV-2 VUI 202012/01” (i.e., variant under investigation from December 2020). Terms such as “variant,” “strain,” and “lineage” are used by the press and the scientific community interchangeably. The variant is also often called B.1.1.7 from the name of its distinct phylogenetic cluster.
VUI 202012/01 has been identified through viral genome sequencing for the first time in London and the South East of England. It is defined by multiple spike protein mutations (deletion 69-70, deletion 144, N501Y, A570D, D614G, P681H, T716I, S982A, D1118H) as well as mutations in other genomic regions. The most common mutation of this variant, N501Y, is on the spike protein responsible for infecting human cells.
Why is this variant more transmissible?
It is common to see multiple variants of the same virus; that is why we have a different flu vaccine every year. New strains do not mean that the virus is more harmful, and there is currently no evidence that this variant causes more severe disease or higher mortality. We also know that mortality is a lagging indicator, which will need to be assessed over the coming weeks. However, current evidence shows that infection rates in geographical areas where the VUI 202012/01 has been circulating have increased faster than expected, and the modelling evidence has demonstrated that this variant has a higher transmission rate than the variants in current circulation. The reasons why this is happening can be multiple.
B.1.1.7 has an unusually large number of mutations, particularly on the spike protein. Three of these mutations have effects that could potentially explain the higher rate of transmission of the variant:
- Mutation N501Y occurs in one of the six key residues within the receptor-binding domain (RBD) and has been described as increasing binding affinity to human and murine ACE2 receptor (the “entry” receptor for the virus).
- The spike protein deletion 69-70 del leads to a conformational change in the spike protein, which has been shown to increase the evasion to the human immune response.
- Mutation P681H is adjacent to the S1/S2 furin cleavage site, a site with high variability in coronaviruses.
This overall increased affinity of the new variant for the human cells means that if you breathe in the new COVID-19 variant, you are more likely to be infected because it binds more strongly to the cells. Additionally, it also appears that this strain is better at evading the immune system, meaning that the standard defence mechanisms will not be as efficient.
Is this new strain affecting the detection of the virus?
Some diagnostic laboratories in the UK have reported that the deletion 69-70del in the spike protein causes negative results from RT-PCR assays for S-gene. However, assays targeting this specific gene are not widely used for primary detection. Only one assay targeting the S-gene is on the list of published assays listed by the WHO. Since mutations are more likely to occur in this gene, relying only on the S-gene for primary detection of SARS-CoV-2 infection is not recommended.
Will vaccines protect from this variant?
There is currently no evidence that suggests that the current vaccines being deployed would not protect the population against the new strain. Scientists say that a change in vaccine effectiveness is unlikely. However, there are no data available yet with respect to the ability of antibodies elicited by vaccines under development to neutralize this variant. Further laboratory research is currently being undertaken to obtain a detailed phenotypical characterization of the new variant, and results are expected in the coming weeks.